Introduction
Coronaviruses infect the human population causing moderate respiratory diseases, but more severe cases frequently occur that can even be fatal. SARS-CoV-2 is one of seven viruses from the Coronaviridae family that infect humans, four of which (229E, OC43, NL63, and HKU1) cause seasonal diseases with symptoms similar to those of the common cold1,2. Other recently discovered viruses in the family are the one that causes severe acute respiratory syndrome (SARS-CoV) and the coronavirus that causes Middle East respiratory syndrome (MERS-CoV), which are of zoonotic origin and have been associated with acute and fatal diseases, severely compromising the respiratory system and, therefore, the lives of the hosts3.
Coronaviruses have caused outbreaks or pandemics, such as that caused in 2002-2003 by SARS-CoV, infecting 8,437 people with a mortality rate of 9.6%4. In 2012, MERS-CoV infected 2,494 people, with a mortality rate of 34.4%5. Finally, the recent pandemic in 2019-2020 caused by SARS-CoV-2 led to the disease called COVID-19 (Coronavirus Disease 2019), and to date, 700 million confirmed cases and approximately 7 million deaths have been documented6. The COVID-19 pandemic began in December 2019 in the city of Wuhan, Hubei Province, China, in a group of people who presented pneumonia of unknown origin, suspected of having been infected in a market where animal foods were sold7.
Biology of SARS-CoV-2
Coronaviruses are enveloped viruses with a positive-sense RNA genome. The genome size is 30 to 32 kb and they belong to the order Nidovirales, family Coronaviridae, subfamily Coronavirinae2. Within this subfamily, there are four different genera: Alphacoronavirus, Betacoronavirus, Gammacoronavirus, and Deltacoronavirus; SARS-CoV-2 belongs to the Betacoronaviruses7. The nucleocapsid protein forms complexes with the genomic RNA to form a helical capsid structure that is found within the viral envelope. The spicule or Spike (S) protein is found in the form of trimers, which form structures called peplomers that are embedded in the virion envelope, giving it the shape of a crown. In certain coronavirus virions, the envelope protein (E) forms smaller spikes on the membrane8 (Fig. 1).
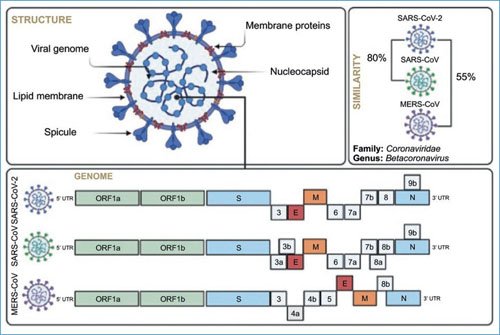
Figure 1. Viral biology: structure, proteins, and genome of SARS-CoV-2, the etiological agent of COVID-19. Comparison of the genome structure with other closest human betacoronaviruses (SARS-CoV and MERS-CoV), and nucleotide homology.
The 5’ end of the coronavirus genome encodes the replicase gene, which contains two open reading frames (ORFs), ORF1a and ORF1ab, spanning approximately 20 kb, about two-thirds of the genome (Fig. 1). ORF 1a and 1ab encode 16 non-structural proteins (NSP). Many of these NSPs have enzymatic activities, including proteases, RNA-modifying enzymes, as well as polymerases and helicases. The 3’ end is the region that encodes the structural proteins: membrane (M), envelope (E), nucleocapsid (N), and Spike (S). Additionally, at the 3’ end is also the region that codes for accessory proteins 3, 6, 7a, 7b, 8, and 9b, whose function remains unclear, although it is known that they may have functions to inhibit the innate immune response9.
Replication cycle
Infection of the target cell begins with binding to the angiotensin-converting enzyme 2 (ACE2) receptor and proteolytic cleavage of the S protein, either extracellularly by transmembrane serine protease-2 or by furin, in cytoplasmic vesicles (Fig. 2). Proteolytic cleavage of the S protein results in the formation of two subunits S1 and S2; in the latter is the fusion peptide that interacts with the cell membrane, resulting in the final entry of viral genomic RNA into the cytoplasm10. Because the genomic RNA is positive-sense, it can be taken as a template by cellular ribosomes for the production of the ORF1a and ORF1ab polypeptides. The first proteins synthesized from these polypeptides form a replication complex and are mainly enzymes (RNA-dependent RNA polymerase) responsible for transcription and production of subgenomic and genomic RNA. Subgenomic RNAs encode the structural proteins (N, E, M, and S) and accessory proteins. The genomic RNA is assembled with nucleocapsid proteins and viral proteins anchored in cell membranes, forming new virions that are released from the cell through budding11 (Fig. 2).
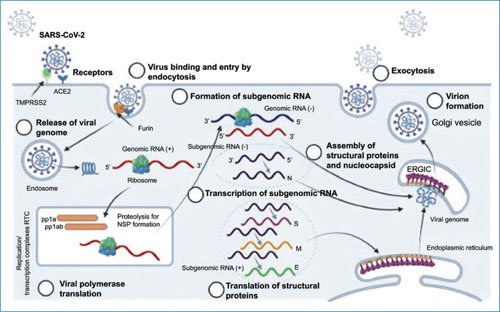
Figure 2. SARS-CoV-2 replication cycle. Recognition of the cellular receptor, entry and fusion of cellular and viral membranes for the subsequent entry of the viral genome into the cytoplasm. The cycle is completed with viral transcription and protein synthesis for the assembly of new viral particles and their release by budding. ERGIC: endoplasmic reticulum-Golgi intermediate compartment; NSP: non-structural proteins.
Origin
To date, there is insufficient evidence to indicate when and where this new virus emerged, but the most accepted theory is that of zoonotic transmission from bat through an intermediate host and finally to humans. This hypothesis is based mainly on the analysis of nucleotide sequences of other Betacoronaviruses present in some mammals, such as rodents and bats7. The Betacoronaviruses with the highest nucleotide homology are those found in bats of the species Rhinolophus affinis and Rhinolophus sinicus, which show 96% and 86% homology, respectively. Another Betacoronavirus very similar in sequence is the one present in the mammal Manis javanica (pangolin), which has 91% homology with SARS-CoV-2; however, it shows more than 95% homology (Fig. 3) in the receptor-binding domain (RBD) region12. These three species could be in frequent contact due to illegal trafficking of live species for subsequent human consumption in China. Their interaction could be the mechanism by which bat viruses infect other mammals that can function as intermediate hosts to finally infect humans.
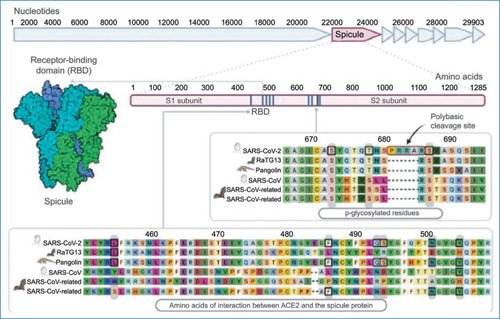
Figure 3. Spike (S) protein of SARS-CoV-2 and receptor-binding domain (RBD). The S protein has approximately 1,300 amino acids and in the S1 region is the RBD; at least six positions of interaction with the ACE2 cellular receptor are known. In the S2 region is the polybasic cleavage site (boxes). An alignment of the amino acid sequence of these regions is shown comparing the SARS-CoV-2 sequence with that of other betacoronaviruses from birds and other mammals. ACE2: angiotensin-converting enzyme 2.
The Spike protein of SARS-CoV-2
The S protein is the major surface antigen of the virus and its main function is to bind to the cellular receptor and mediate the membrane fusion required for cell entry and initiation of infection; therefore, it determines the host range and viral tropism13. Because of this, the S protein is the primary target of neutralizing antibodies generated during infection and also a key focus in vaccine design.
The S protein is a trimeric protein and each monomer is synthesized as a single polypeptide chain of approximately 1,300 amino acids. In many coronaviruses, the S protein is processed by host proteases generating two functional subunits (S1 and S2) (Fig. 3) that remain non-covalently linked in the prefusion conformation11. These structures are important for cellular receptor recognition, virus entry, and fusion10. The S1 subunit contains the RBD region, which includes amino acids 357 to 528. Within this region, there are at least six positions that are essential for binding to the cellular receptor: 445, 486, 493, 494, 501, and 505 (Fig. 3). Adjacent to these positions are amino acids that are important for antigen presentation and stimulation of humoral and cellular immune responses13.
ACE2 cellular receptor and viral tropism
The ACE2 protein is a member of a family that has wide distribution in various tissues. In terms of its function, ACE2 has diverse biological functions, including the regulation of blood pressure through the renin-angiotensin-aldosterone system, which is linked to the pathogenesis of cardiovascular diseases14. Particularly in the pancreas, it plays an important role as a protector of glycemia, while in the kidney, low levels of ACE2 are associated with progressive diseases such as diabetic nephropathy14,15.
One of the main characteristics of COVID-19 is the large number of organs that are affected, largely because many organs express the ACE2 protein. Histopathological data have shown that the virus has tropism for different cell lineages and organs, such as the renal, circulatory, and neurological systems, as well as pharyngeal and gastrointestinal tissue, and mainly the respiratory tract16. It is important to mention that after the onset of SARS-CoV-2 infection, secondary effects arise in the pathophysiology of various organs, as well as direct viral toxicity, damaging endothelial cells, causing thromboinflammation and dysregulation of the renin-angiotensin-aldosterone system. It should be noted that the alteration in the renin-angiotensin- aldosterone system is believed to be exclusive to COVID-1916. Likewise, immune pathogenesis caused by overproduction and release of proinflammatory cytokines that significantly affect the lungs, as well as dysregulation in microcirculation, could also be a result of viral sepsis17. SARS-CoV-2 infection in humans is possible due to the existence of specific host cellular receptors (SARS-CoV, SARS-CoV-2, and HCoV-NL63 recognize ACE2, MERS-CoV recognizes dipeptidyl peptidase 4, and OC43 and HKU1 recognize 5-N-acetyl-9-O-acetyl-sialidase) that allow virus entry into the cell, to subsequently complete its cycle, and spread effectively in the human population18. On the other hand, it has been described that ACE2 polymorphisms in human cells could modify the interaction with SARS-CoV-2, increasing or decreasing the affinity of ACE2 with the RBD of the virus S protein and, in some cases, favoring viral infection19.
Natural evolution of the disease
The infection begins with direct and indirect exposure to a source of infection, which allows the virus to enter the respiratory system, and interact with the ACE2 receptor present in epithelial cells20 (Fig. 4). Since there is a wide distribution of the ACE2 receptor in the upper respiratory tract, establishment of infection is very likely, associated with a high viral load at the time of exposure21. The main respiratory symptoms begin to appear between 5 and 7 days after exposure. From the establishment of infection in the upper respiratory tract and the oropharyngeal cavity, it spreads to different organs and becomes a systemic infection. Some non-epithelial cells of the mucosa, trachea, and bronchi also express the ACE2 receptor, including type II alveolar cells found in the lung parenchyma, indicating that this receptor is present not only in epithelial cells but also in various parts of the respiratory tract. Therefore, the tracheal and bronchial epithelium is also susceptible to SARS-CoV-2 infection. It should be noted that these primary sites of infection could facilitate the spread of the virus to the alveoli, where the main damage occurs and where ACE2 expression can be stimulated by environmental processes, such as microbial infections or inflammatory processes in the lung20. The first immune response against the virus is the innate response, composed mainly of interferons, and begins before or during the onset of symptoms (Fig. 4). This response is essential for restriction of viral replication in the early stage and for limiting infection to its less severe forms22. Subsequently, the adaptive immune response appears, composed mainly of B lymphocytes, and the production of neutralizing antibodies and T lymphocytes with the cellular response23. The neutralizing antibody response appears to last more than 1 year, and its magnitude depends primarily on exposure to viral antigens, with patients with severe disease having a greater quantity of these antibodies compared with patients who are asymptomatic or have moderate symptoms24.
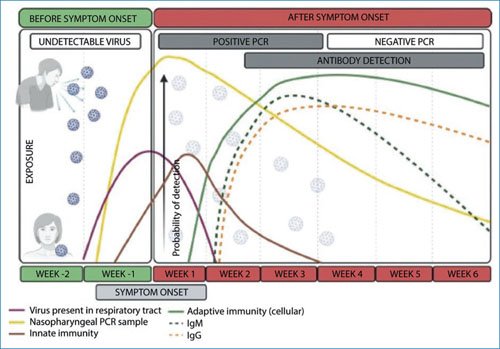
Figure 4. Natural evolution of SARS-CoV-2 infection. After exposure to the virus, infection is established in the upper respiratory tract, which begins with respiratory symptoms between 5 and 7 days after exposure. The virus can be detected before the onset of symptoms in the nasopharyngeal and oropharyngeal cavity, and continues to be shed for at least 3 more weeks. Immune responses against the virus, innate, antibody, and cellular types appear after the onset of symptoms. IgG: immunoglobulin G; IgM: immunoglobulin M; PCR: polymerase chain reaction.
Has SARS-CoV-2 changed since its appearance in Wuhan?
Due to its nature as an RNA genome virus, SARS-CoV-2 has a high mutation rate25, although this is lower compared with other RNA viruses, such as influenza26. Several factors are involved in this biological variation: high transmission in the population, host factors, and the use of vaccines can increase it, and therefore, favor the appearance of changes or mutations throughout the viral genome27. Since its emergence at the end of 2019, most proteins have undergone changes, mainly NSP3, NSP12b, ORF3a, N, NSP2, NSP4, ORF8, NSP14, NSP13 and, importantly, the S protein28.
The first notable change was at site 614 (D614G), which was initially detected in England and spread rapidly through Europe and North America. This change defined the appearance of new viral clades (G, GH, GV, GR, GK), which predominated in Europe and North America from March 2020. In vitro and epidemiological studies indicate that this change conferred increased transmissibility and pathogenicity to the virus29. The change at position 614 occurs in the interaction zone of the S protein monomers, enhancing the ability to adopt an open conformation, which has greater affinity for the cellular receptor30. From September 2020, other changes in the S protein began to be detected in different parts of the world, such as N501Y, E484K, K417N, L452R, and P681H28,30. These mutations are found in regions such as the RBD, in monomer-monomer interfaces and in the cellular furin cleavage site, as well as in immunogenic sites30. It is important to conduct molecular surveillance to detect these mutations in SARS-CoV-2 viruses locally, regionally, and globally.
A viral variant has two or more mutations that can confer a phenotype of greater transmission, immune response evasion, or greater pathogenicity. The first reported variant was the so-called English variant (20I/501Y.V1), which in the PANGO nomenclature (Phylogenetic Assignment of Named Global Outbreak Lineages) is identified as B.1.1.7 and which presents, among others, the N501Y mutation31. The mutation at position 501 alters binding to the cellular receptor, increasing SARS-CoV-2 transmission. To date, this has been confirmed with various studies using strategies ranging from in silico modeling to meta-analyses of data.
Another mutation that has been found recurrently in different parts of the world is the change from glutamic acid to lysine or asparagine at position 484 (E484K or E484Q). This change does not directly affect binding to the cellular receptor; recent studies indicate that it is an important site for recognition by neutralizing antibodies, and the emergence of viruses with this change would imply evasion of the immune response32. Other mutations that would affect the immunogenicity of the S protein and that are emerging in different parts of the world are K417T, L452R, N439K, and Y453F28. Deletions in the S protein that affect its structure have also been detected. These deletions have been found mainly in two positions: 69-70 (Δ69-70) and 144 (Δ144)31. In particular, it has been described that these deletions generate a decrease in binding to neutralizing antibodies that are generated in patients treated with plasma from convalescent patients and from recovered patients who have some type of immunodeficiency33. The emergence of these deletions in these patient groups could suggest that they are being positively selected, and could be a mechanism of resistance to antibodies against SARS-CoV-230. The sites where these deletions are found have been called recurrent deletion regions, and although substitutions are relatively infrequent, they could represent a mechanism driving rapid evolution, which could promote genetic drift34.
Viral variants
Currently, it is possible to classify SARS-CoV-2 by detecting mutations in its genetic sequence. Based on mutations found in populations around the world, its variants can be determined35. A variant is a lineage that has a demonstrably different phenotype from others circulating around the world, with two or more mutations that confer benefits to the virus, such as antigenicity, transmissibility, or virulence36. Variants are classified into variant of interest (VOI), variant of concern (VOC), and variant of high consequence (VHC)36.
At the end of 2020, various SARS-CoV-2 variants began to be detected circulating in different populations around the world. The main VOCs are those found in England (B.1.1.7 or alpha), Brazil (P1 or gamma), South Africa (B.1.351 or beta and B.1.529 or gamma), and India (B.1.617 or delta)31,37-40; Table 1 shows the mutations present in each of these variants. Importantly, it should be noted that some of these variants share mutations, primarily mutations N501Y, E484K, L452R, and P681H35. According to this, it is vital to understand the effects of the presence of these variants on transmissibility, pathogenicity, and immunogenicity, as well as on the effectiveness of vaccines currently being used.
Table 1. Impact on transmission, pathogenicity, and immunogenicity of SARS-CoV-2 variants
| Variant (mutations in Spike) |
Mutations in other genes | Transmission | Pathogenicity | Immunogenicity |
|---|---|---|---|---|
| B.1.1.7/alpha (del 69-70, del 144, N501Y, P681H) | ORF1ab:T1001I ORF1ab:A1708D ORF1ab:I2230T del I1288-9 ORF8:Q27 ORF8:R52I ORF8:Y73C N:D3L N:S235F |
50% increase (A) | Potential increase in severity, based on hospitalization data and fatality rates |
Minimal impact on neutralization by sera from convalescent and vaccinated patients (B) |
| P1/beta (E484K, N501Y, P681H) | ORF1ab: S1188L ORF1ab:K1795Q del 11288:9 ORF3a:G174C ORF8:E92K N:P80R |
No data | No data | Considerable reduction in neutralization by sera from convalescent and vaccinated patients (C) |
| B.1.1.351/gamma (E484K, N501Y) | E:P71L N:T205I ORF1a:K1655N |
50% increase (D) | No data | Considerable reduction in neutralization by sera from convalescent and vaccinated patients (C) |
| B.1.529/omicron (30 mutations in S and 15 in RBD) | 25 mutations distributed throughout the viral genome |
50% increase (E) | Reduction in severity |
Considerable reduction in neutralization of sera from convalescent and vaccinated patients (E) |
| B.1.617.1/delta (T95I, L452R, E484K, P681R) | ORF1ab:T1567I, T3646A, P4715L ORF1ab:G5530C ORF1ab:M5753I ORF1ab:K6711R ORF1ab:S6713A ORF3a:S26L M:I82S, N:R03M, Q377Y, ORF7a, V82A |
No data | No data | Considerable reduction in neutralization by sera from convalescent and vaccinated patients (F) |
Viral variants according to the PANGO/WHO nomenclature and their main mutations in the Spike protein.
In summary, it is known that the alpha variant showed greater transmissibility, while the beta, gamma, and delta variants, in addition to having greater transmission capacity, escaped sera from convalescent people. Furthermore, it has also been seen that some vaccines, such as the one developed by AstraZeneca, present lower levels of protection against these variants (Table 1). The last VOC emerged at the end of 2021 (omicron) and has at least 30 mutations in the S protein and 15 in the RBD. These mutations confer a greater affinity for the ACE2 receptor and greater evasion of the immune response stimulated by vaccination and by infections with other SARS-CoV-2 variants. In Latin America, new variants different from that found in Brazil were detected. In Colombia, since the end of 2019, a variant (B.1.1.621 or mu) was detected that shares several mutations with other VOCs, such as mutations E484K, N501Y, and P681Q41. Despite having these changes, it is still not considered a VOI or VOC. Another variant of interest, already accepted by the World Health Organization (WHO), is one the lambda variant, which was first detected in Peru and presents mutations L452Q and F450S in S36. On the other hand, in Mexico, our working group and other institutions in the country detected the presence of a new lineage (B.1.1.519) since late December 2020, which increased its presence during the months of January to March 202142. This lineage presents two important mutations, T478K and P681H, for which it was suggested to propose it as a VOI42. This lineage emerged at the end of 2020 in central Mexico and also presents other mutations in other viral genes: N:G204R, N:R203K, ORF1a_P959S, ORF1a_I3618V, ORF1a_T3255I, and ORF1a_T4175I.
The VOC omicron, which emerged at the end of 2021, has remained since then and diversified into various lineages and sublineages, with the latest to appear being those derived from sublineage BA.2.86.1 or JN.1. The omicron variant, originally designated B.1.1.529, diversified into two branches in BA.1 and BA.2, and the latter in turn diversified into BA.4 and BA.5 in early 2022. These branches diversified over time, with the predominant subvariants being BQ derived from BA.2. On the other hand, recombination occurred in two BA.2 sublineages, BJ.1 and BM.1.1.1, and in April 2023, the XBB subvariant was detected, with XBB.1.5 being predominant for several months worldwide43. In August 2023, the WHO designated subvariant BA.2.86 as a variant under monitoring because it has 33 mutations in S and 14 in RBD, compared with its predecessor BA.2. From this subvariant, JN.1 emerged, and at the end of 2024, variants KP, XEC, and LP were detected, the latter dominant in 202544.
Global and Mexican molecular surveillance
What we know about SARS-CoV-2 from a molecular point of view is due to knowledge of viral genomes present in clinical samples. This is done through next- generation sequencing and bioinformatics methods. With these methodologies, it has been possible to identify the etiological agent causing this new disease and it has been possible to identify viral evolution for more than 1 year since its emergence, as well as the emergence of new variants that can affect the course of the disease and vaccination. In Mexico, different health institutions, including the National Institute of Respiratory Diseases Ismael Cosío Villegas, as well as academic institutions, have conducted this molecular surveillance, and by June 2021, more than 10,000 complete genomes from most of the states of the Republic had been deposited in the GISAID database. These data are necessary to better understand the pandemic in our country, as well as to generate knowledge that helps us address the health emergency in the coming months and years. Likewise, this knowledge can also help us understand virus evolution and its adaptation to humans.
Vaccination and disease control perspectives
The pandemic caused by SARS-CoV-2 has been the driving force for the development of new vaccination strategies, such as those based on RNA, and the use of others that were beginning to be used, such as adenovirus vectors. In addition, several strategies already known with other vaccines have been used, such as the use of peptides, recombinant proteins, and inactivated viruses, among others (Fig. 5)45.
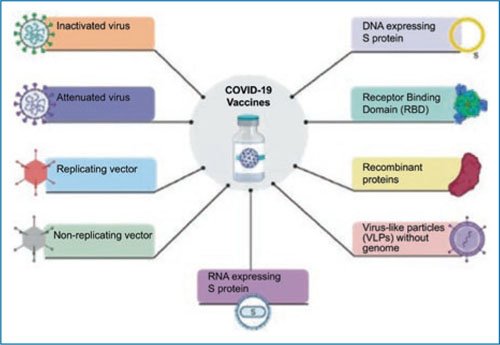
Figure 5. Repertoire of vaccines against SARS-CoV-2 developed using various strategies; among the most successful, the use of RNA in nanoparticles by BioNTech-Pfizer and adenovirus vectors by the Oxford-AstraZeneca consortium, the Gamaleya Institute, and the Cansino company.
Several academic institutions from different countries have formed alliances with large pharmaceutical companies for the mass development of vaccines. Currently, the most widely used vaccines worldwide are the one developed by BioNTech and Pfizer using RNA, the Oxford-AstraZeneca vaccine, the vaccines from the Gamaleya Institute in Russia and the Beijing Institute43, the Cansino vaccine, which uses adenovirus vectors, and Sinovac’s inactivated vaccine. Of these, the vaccine that offers the greatest protection is Pfizer’s, close to 90%, while Cansino has the lowest protection (50-60%)46,47. The emergence of new variants has been a challenging situation for vaccine efficiency. mRNA-based vaccines have included variants XBB.1.5 and JN.1 in their new formulations as an update of the S protein. Although these new vaccines stimulate neutralizing antibodies against the new variants, there are insufficient studies to know if there is an advantage regarding the reduction of severe cases of the disease48.
In the absence of easily accessible antiviral drugs, vaccination is currently the most effective control measure, which in the medium and long term will result in a reduction in active cases, hospitalizations, and deaths. Molecular characterization of SARS-CoV-2 is essential to monitor the emergence of mutations in the virus that may affect vaccine effectiveness or lead to the emergence of more pathogenic variants49.
Funding
None.
Conflicts of interests
The authors declare that they have no conflicts of interests.
Ethical considerations
Protection of human and animal subjects. The authors declare that no experiments were performed on humans or animals for this research.
Confidentiality, informed consent, and ethical approval. The study does not involve personal patient data or require ethical approval. SAGER guidelines do not apply.
Statement on the use of artificial intelligence. The authors declare that they did not use any type of generative artificial intelligence for the writing of this manuscript.